Tidyverse Other packages
tidyr - Data Tidying
A package in the Tidyverse for tidying data
Helps convert data between wide and long formats
Simplifies reshaping and cleaning datasets
Key Functions in tidyr
pivot_longer()
Converting from Wide data (multiple columns for the same variable) to Long data (one column for each variable)
# A tibble: 6 × 3
id year value
<int> <chr> <dbl>
1 1 2021 100
2 1 2022 110
3 2 2021 150
4 2 2022 160
5 3 2021 200
6 3 2022 210pivot_wider()
Converting from Long to Wide Format
# A tibble: 3 × 3
id `2021` `2022`
<dbl> <dbl> <dbl>
1 1 100 110
2 2 150 160
3 3 200 210separate()
Splits a single column into multiple columns
# A tibble: 3 × 2
first_name last_name
<chr> <chr>
1 John Doe
2 Jane Smith
3 Alice Johnson unite()
Combines multiple columns into one
# A tibble: 3 × 1
full_name
<chr>
1 John Doe
2 Jane Smith
3 Alice Johnsondrop_na()
Removes rows with missing values
# A tibble: 1 × 2
x y
<dbl> <dbl>
1 1 4fill()
Fills missing values using the previous or next available value
# A tibble: 5 × 1
x
<dbl>
1 1
2 1
3 3
4 3
5 5tidyr vs dplyr
tidyr: Specializes in reshaping and tidying data (e.g., pivot_longer(), pivot_wider(), separate()).
dplyr: Specializes in data manipulation, such as subsetting, grouping, and summarizing (e.g., filter(), mutate(), summarize()).
readr - Data Import/Export
part of the tidyverse, focused on reading and writing rectangular data (e.g., CSV, TSV).
Faster and more consistent than base R functions
Makes data handling easy and efficient.
Key Functions:
read_csv()
# A tibble: 5 × 1
x
<dbl>
1 1
2 NA
3 3
4 NA
5 5read_delim()
# A tibble: 5 × 1
x
<dbl>
1 1
2 NA
3 3
4 NA
5 5write_csv()
write_delim()
purrr - Functional Programming
enhances R’s functional programming capabilities by providing a consistent, simple way to apply functions to lists, vectors, and other data structures.
Key function:-
map()
Applies a function to each element of a list and returns a list.
[[1]]
[1] 1
[[2]]
[1] 4
[[3]]
[1] 9
[[4]]
[1] 16
[[5]]
[1] 25map_dbl()
To apply a function and return a vector (not a list), map_dbl() is used.
[1] 1 4 9 16 25map2()
Applies a function to two inputs simultaneously.
useful when you need to operate on two lists.
[[1]]
[1] 11
[[2]]
[1] 22
[[3]]
[1] 33map_chr()
Used to get a character vector instead of a numeric one.
[1] "1" "2" "3" "4"purrr vs tidyr
purrr is specifically focused on working with lists and vectors using functional programming techniques.
tidyr is focused on reshaping data frames (long to wide format and vice versa), separating and combining columns.
tibble - Modern Data Frames
A modern version of a data frame.
More robust and user-friendly than traditional data frames.
Improved handling of large data sets
Basic Tibble
# A tibble: 3 × 3
Name Age Location
<chr> <dbl> <chr>
1 John 25 New York
2 Alice 30 London
3 Bob 22 Paris Tibble with Mixed Data Types
# A tibble: 3 × 4
Name Age Is_Active Height
<chr> <dbl> <lgl> <dbl>
1 John 25 TRUE 5.9
2 Alice 30 FALSE 5.6
3 Bob 22 TRUE 6.1Tibble vs. Data Frame
Data Frame
- Displays all rows, which can be overwhelming with large datasets.
- Automatically converts character vectors to factors (unless specified otherwise).
Tibble
Only shows the first few rows, making it easier to handle large datasets.
Does not convert character columns into factors by default, avoiding a common issue with data frames.
stringr - String manipulation
Provides simple functions for common text operations.
Focuses on consistency and ease of use.
Key Functions:-
str_detect()
[1] TRUEstr_replace()
[1] "I love Python programming."str_replace_all()
[1] "The dog is on the mat. The dog is cute."str_sub()
[1] "Hello"str_to_lower()
[1] "hello, world!"str_split()
[[1]]
[1] "apple" "banana" "cherry"str_length()
[1] 5lubridate
It makes working with date-times easier
Provides functions to manipulate, parse, and format date-time data.
Simplifies operations like extracting parts of date-time, arithmetic operations, and handling time zones.
Parsing Dates
[1] "2025-03-03"Parsing Dates with Time
[1] "2025-03-03 12:30:45 UTC"Extracting Date-Time Components
[1] 2025[1] 3[1] 3[1] 12[1] 30[1] 45Handling Time Intervals
[1] 2025-03-01 UTC--2025-03-05 23:59:59 UTCforcats - Working with Factors
Makes working with categorical variables easier.
Provides tools to manipulate factors and handle tasks like reordering, renaming, and combining levels.
Key functions :
Creating Factors
[1] Low High Medium Low High
Levels: High Low MediumReordering factor levels
[1] Low High Medium Low High
Levels: Medium High LowChanging Factor Levels
[1] Very Low Very High Medium Very Low Very High
Levels: Very High Very Low MediumCombining Factor Levels
[1] Low/Medium High Low/Medium Low/Medium High
Levels: High Low/MediumWorking with Factor Levels
[1] Low High Medium Low High
Levels: High Low MediumVisualizing Factor Data
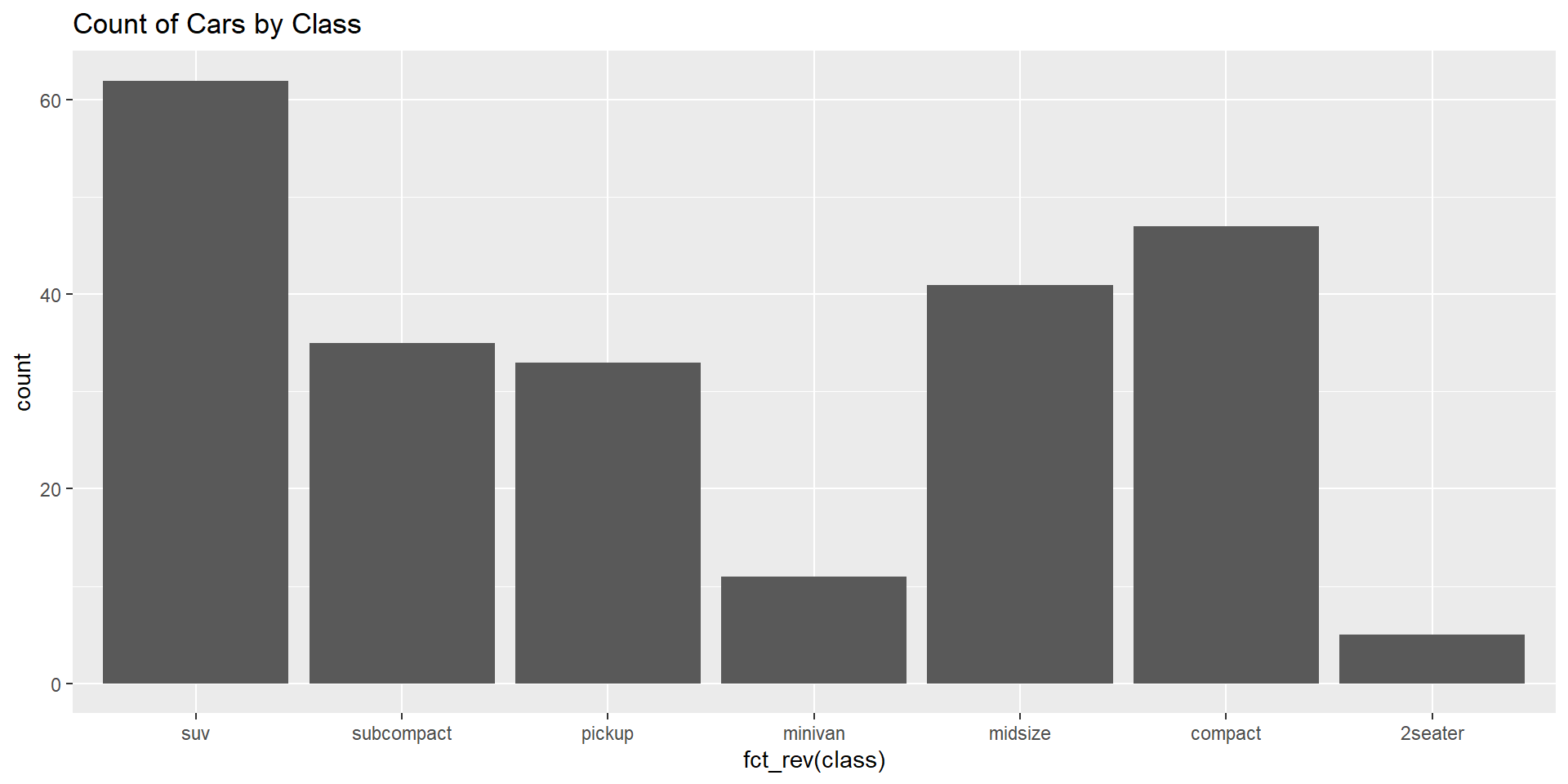
forcats vs dplyr
forcats works with factors used for modifying factor levels, whereas dplyr also provides some functions that manipulate factors (e.g., mutate() and factor()).
forcats: Designed specifically for working with factors and making factor level manipulations easier.
dplyr: While dplyr can work with factors, forcats is more specialized for factor-specific operations.